Software development
Recent developments in experimental techniques and computer simulations provided the basis to achieve many of the breakthroughs in understanding materials down to the atomic scale. While extremely powerful, these techniques produce more and more complex data, forcing all departments to develop advanced data management and analysis tools as well as investing into software engineering expertise.
As a consequence, a MPIE software task force has been established, where software engineering and simulation software expertise developed e.g. in the CM department is utilized. Here, we have selected a few examples to demonstrate how these developments help to propel science both in experiment and theory.
An integrated development environment
Pyiron has been introduced as an integrated platform for materials simulations and data management. At the same time, pyiron has also modified the culture of research in the CM department towards a stronger focus on software development. For the outer community, this is visible by a growing number of journal publications that are combined with executable code required for the physical analysis shown in the paper [1]. This code is provided within binder containers, such that the user can directly employ the simulations on cloud resources without the need of cumbersome installations on the local hardware. One example of this kind is the fully automated approach to calculate the melting temperature of elemental crystals [2]. The manuscript explains many of the technical challenges connected to the interface method, used for melting point calculations. The underlying complex simulation protocols are provided as pyiron notebooks, which allows a much more interactive access, reproduction and exploration of the underlying concepts than a written manuscript.
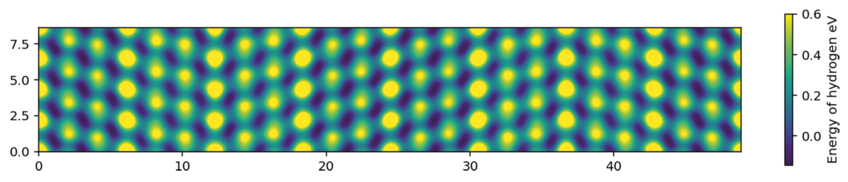
More significant for the research inside the CM department is, however, the cooperative development of new physical concepts in regular coding sessions. Several teams are currently meeting for several hours on a weekly basis, in order to test new physical ideas by directly implanting and executing them in pyiron notebooks. The physically intuitive structure of the pyiron environment allows such a research approach, without spending time on many command lines, taxonomy and syntax. For example, one group is exploring the physical nature of thermo desorption spectra (TDS) for hydrogen in metals. This involves the automatic detection of favourable sites for H incorporation into complex microstructure features (such as grain boundaries, Fig. 1), the analysis of possible jump processes between these sites and the evaluation of the H dynamics by kinetic Monte-Carlo simulations. Another group is connecting the workflows in pyiron notebooks with the semantic description of materials knowledge in ontologies, opening new routes of software development based on physical concepts rather than command lines.

Machine learning for solid mechanics
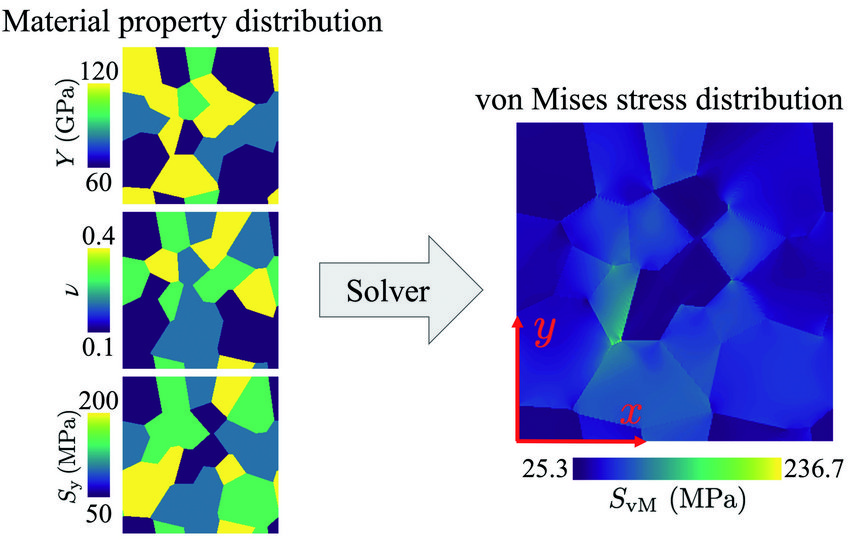
In the MA department, an artificial intelligence (AI)-based solution for predicting the stress field in inhomogeneous elasto-plastic materials was recently developed [3]. The training data for a convolutional neural network (CNN) were generated using DAMASK (Düsseldorf Advanced Material Simulation Kit) for modelling multi-physics crystal plasticity. Both, cases of materials with elastic and elasto-plastic behaviour, are considered in the training data. The trained CNN model is able to predict the von Mises stress fields (Fig. 2) orders of magnitude faster than conventional solvers. In the case of elasto-plastic material behaviour, conventional solvers rely on an iterative solution scheme for the nonlinear problem. The AI-based solver is able to predict the correct solution in one step forward evaluating of the CNN. This significantly reduces the implementation complexity and computation time. The augmentation of such a fast stress estimator with finite element-based solvers will open new avenues for the multi-scale simulation of materials.
Software tools for atom probe microscopists
In order to achieve scientific conclusions from atom probe tomography (APT), these data have to be processed into tomographic reconstructions, i.e. 3D models that specify the approximate original locations of the atoms in the field-evaporated specimen. Representing point clouds, these models are post-processed with methods from the field of spatial statistics, computational geometry, and AI resulting in quantified (micro)structural features or patterns that can be related to descriptors in physics-based models at the continuum scale such as dislocations, grain and phase boundaries, or precipitates.

It is important to quantify the eventual uncertainty and sensitivity of the descriptors as a function of the parameterization. To this end, though, the employed proprietary tools face substantial practical and methodological limitations with respect to the accessibility of intermediate (numerical and geometry) data and metadata they offer, the transparency of the algorithms, or the functionalities they provide to script and interface proprietary tools with open-source software of the atom probe community.
Involving APT domain specialists from the MA and computational materials scientists from the CM department, we implemented a toolbox of complementary open-source software for APT, which makes efficient algorithms of the spatial statistics, computational geometry, and crystallography community accessible to support scientists with quantifying the parameter sensitivity of APT-specific methods they use. Our tools exemplify how open-source software enables to build tools with programmatically customizable automation and high-throughput characterization capabilities, whose performance can be augmented by cluster computing. We show how to automate and make exactly reproducible the execution of microstructural feature characterization [4 - 6] for atom probe data.
Analysis tools for complex imaging and spectroscopy
One example of software development in the SN department is the creation of analysis workflows for data from transmission electron microscopes (TEM). Typically, most experimentalists use graphical user interface (GUI) based tools to process their data. Many of these tools are proprietary and licensed by the hardware manufacturer. The analysis options in this software are limited and rarely reproducible. In addition, with the advancement in detector technology and other instrumentation, the size and complexity of data is ever increasing. This calls for highly customizable analysis tools. This is also necessary for bridging the gap between the powerful machine learning and artificial intelligence libraries in the scientific python ecosystem and the workflow of the average experimentalist.
The experimental TEMMETA [7] library was built to address the most common analysis needs offered in software like ImageJ or Digital Micrograph, such as drawing line profiles, selecting subsets of the data, visualizing the data, and calculating strain fields from atomically resolved images. TEMMETA also has the feature of automatically storing information about each operation performed on data in a history attribute, thereby making it easier to reproduce the analysis. As development progressed, it became clear that the TEMMETA project had many of the same aims as the Hyperspy project, a library that has existed for a longer time and already has an established community of users and developers. Therefore, it was decided to integrate the novelties of the TEMMETA project into Hyperspy [8].
File parsers to a common representation are essential for bringing the full functionality of Hyperspy to any kind of microscopy data. The SN department is increasingly adopt-ing 4D-STEM as an advanced TEM based technique. It produces large and rich datasets of diffraction patterns collected on a grid of scan points. A typical workflow for these datasets relevant to metallurgists is analysing local crystal orientation via template matching. Existing proprietary software solutions could not handle the data from pixelated detectors, hence a custom built fast and scalable solution [9] was implemented in a package that builds on Hyperspy. This should make the transparent analysis of 4D-STEM datasets more accessible to researchers at the MPIE and all over the world.
References
1. https://pyiron.org/publications/
2. Zhu, L.-F.; Janssen, J.; Ishibashi, S.; Körmann, F.; Grabowski, B.; Neugebauer, J.: Comp. Mater. Sci. 187 (2021) 110065.
3. Mianroodi, J. R., H. Siboni, N. & Raabe, D.: npj Comput. Mater. 7 (2021) 99.
4. Kühbach, M.; Bajaj, P.; Zhao, H.; Celik, M.H.; Jägle, E.A.; Gault, B.: npj Comput. Mat. 7 (2021) 21.
5. Kühbach, M.; Kasemer, M.; Gault, B.; Breen, A.: J. Appl. Cryst. 54 (2021) 1490.
6. Kühbach, M.; London, A.J.; Wang, J.; Schreiber, D.K.; Mendez, M.F.; Ghamarian, I.; Bilal, H.; Ceguerra, A.V.: Microsc. Microanal. (2021). doi: 10.1017/S1431927621012241.
7. Cautaerts, N.; Janssen, J.: Zenodo: din14970/TEMMETA v0.0.6 (2021). doi.org/10.5281/zenodo.5205636.
8. de la Peña, F.; Prestat, E.; Fauske, V.T.; Burdet, P.; Furnival, T.; Jokubauskas, P.; Lähnemann, J.; Nord, M.; Ostasevicius, T.; MacArthur, K.E.; Johnstone, D.N.; Sarahan, M.; Taillon, J.; Aarholt, T.; pquinn-dls; Migunov, V.; Eljarrat, A.; Caron, J.; Poon, T.; Mazzucco, S.; Martineau, B.; actions-user; Somnath, S.; Slater, T.; Francis, C.; Tappy, N.; Walls, M.; Cautaerts, N.; Winkler, F.; Donval, G.: Zenodo: hyperspy v1.6.4 (2021). doi.org/10.5281/zenodo.5082777.
9. Cautaerts, N.; Crout, P.; Ånes, H. W.; Prestat, E.; Jeong, J.; Dehm, G.; Liebscher, C.H.: cond-mat arXiv:2111.07347